Uncategorized
Trump administration’s drug strategy is at odds with recent actions on funding, policy
The White House’s new strategy for addressing the nation’s drug crisis calls for a number of consensus public health measures: the overdose-reversal medication naloxone, medication-assisted treatment, and test strips used to detect fentanyl or other drug supply adulterants.
But the May 4 document appears to run counter to many of the Trump administration’s latest drug policy actions. In particular, it comes just days after the administration issued new restrictions on using federal dollars to distribute test strips and warned against the use of medication-assisted treatment unless accompanied by other services, like counseling.
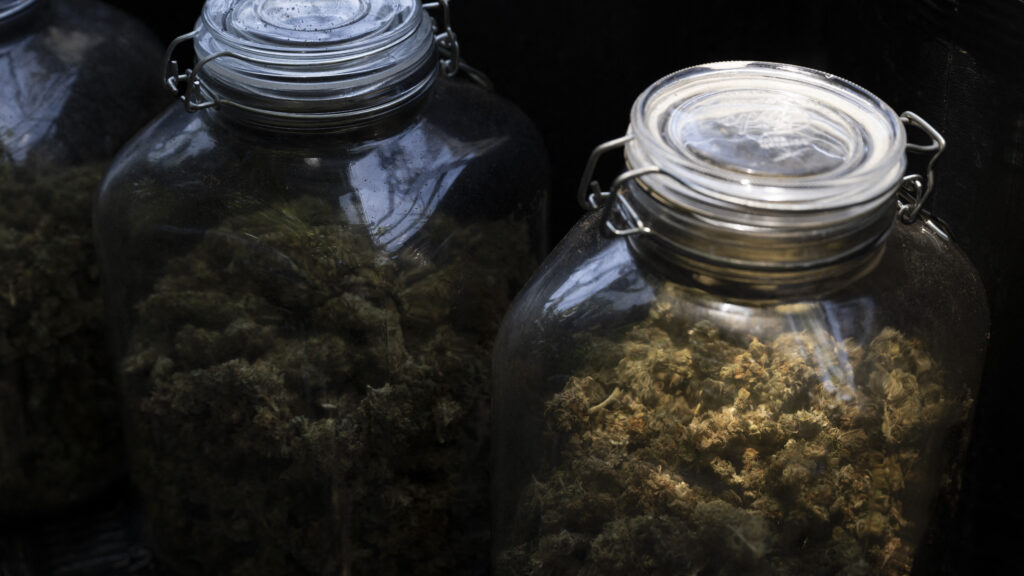

The White House’s new strategy for addressing the nation’s drug crisis calls for a number of consensus public health measures: the overdose-reversal medication naloxone, medication-assisted treatment, and test strips used to detect fentanyl or other drug supply adulterants.
But the May 4 document appears to run counter to many of the Trump administration’s latest drug policy actions. In particular, it comes just days after the administration issued new restrictions on using federal dollars to distribute test strips and warned against the use of medication-assisted treatment unless accompanied by other services, like counseling.
Uncategorized
STAT+: Oregon hospitals won’t outsource to national physician chain after all
After a tidal wave of blowback that culminated in a lawsuit, a nonprofit health system has reversed course in its plan to replace its Oregon emergency physicians with a national chain.
PeaceHealth’s announcement Wednesday didn’t disclose what prompted its change of heart, but those familiar with the situation say it’s because the health system’s plan was poised for defeat in a legal challenge. When PeaceHealth said in February it was cutting ties with Eugene Emergency Physicians, the local group that had staffed its Oregon hospitals for 35 years, the news drew tremendous pushback from doctors, nurses, lawmakers, mayors, and emergency medicine groups.
Then, on March 20, the Eugene emergency physicians sued, arguing that PeaceHealth’s plan to use the Atlanta-based staffing chain ApolloMD violated a new Oregon law prohibiting managed service organizations from directly owning medical practices or interfering with clinical decisions. The case has had four hearings, in which the judge was “quite clear” that the scheme violated the law, Senate Bill 951, said Hayden Rooke-Ley, an attorney who represented the doctors and a senior fellow for health care with the American Economic Liberties Project.

After a tidal wave of blowback that culminated in a lawsuit, a nonprofit health system has reversed course in its plan to replace its Oregon emergency physicians with a national chain.
PeaceHealth’s announcement Wednesday didn’t disclose what prompted its change of heart, but those familiar with the situation say it’s because the health system’s plan was poised for defeat in a legal challenge. When PeaceHealth said in February it was cutting ties with Eugene Emergency Physicians, the local group that had staffed its Oregon hospitals for 35 years, the news drew tremendous pushback from doctors, nurses, lawmakers, mayors, and emergency medicine groups.
Then, on March 20, the Eugene emergency physicians sued, arguing that PeaceHealth’s plan to use the Atlanta-based staffing chain ApolloMD violated a new Oregon law prohibiting managed service organizations from directly owning medical practices or interfering with clinical decisions. The case has had four hearings, in which the judge was “quite clear” that the scheme violated the law, Senate Bill 951, said Hayden Rooke-Ley, an attorney who represented the doctors and a senior fellow for health care with the American Economic Liberties Project.
Uncategorized
Microproteins and Peptideins Expand Boundaries of the Human Proteome
A research team led by scientists at the Princess Máxima Center for Pediatric Oncology, the University of Michigan Medical School, EMBL European Bioinformatics Institute, and the Institute for Systems Biology, has uncovered more than 1,700 new proteins that could have implications for human diseases, including cancer.
Mostly very small, these proteins have been discovered in what’s known as the “dark proteome,” which covers gene products from previously overlooked sections of DNA. These proteins have unusual properties, motivating scientists to coin a new concept, peptideins, to help understand their potentially unique biology. Research co-lead Sebastiaan van Heesch, PhD, a group leader at the Princess Máxima Center, commented, “We know that the current overview of recognized proteins doesn’t capture the full picture. With this study, we show that thousands of overlooked genetic sequences contribute to the dark proteome by producing a new class of protein-like molecules, microproteins, that had been missed before now. But for most of them, we don’t yet know what they do.”
Research co-lead and co-corresponding author Robert Moritz, PhD, professor and head of proteomics at the Institute for Systems Biology, further noted, “Biology has long relied on a relatively small cast of well-characterized proteins to explain the regulatory logic of the cell, but peptideins suggest that beneath that familiar layer lies an entire untapped layer of molecular actors whose functional roles in gene regulation, signaling, and cytopersistence, many we are only beginning to imagine. Given their smaller size and the diversity of cellular contexts in which they appear, I believe peptideins may prove to be among the most versatile and consequential regulatory molecules we have yet encountered in human biology. This is not the end of a search—it is the opening of a vast and fertile new territory for the entire scientific community to explore and exploit, and I look forward to seeing what the broader scientific community uncovers as these molecules, and many more that are yet to be confirmed, are brought into the light.”
Research co-lead John Prensner, MD, pediatric neurooncologist at the University of Michigan Medical School, together with Van Heesch and Moritz, are co-senior and co-corresponding authors of the researchers’ published paper in Nature titled “Expanding the human proteome with microproteins and peptideins.” The team is sharing its discoveries with scientists worldwide in an open-source format to stimulate further research.
Van Heesch added, “With growing interest in industry and academia, peptideins are at the center of multiple drug development initiatives. Similarly, we see them increasingly turning up as important players in diseases, including childhood cancers. We hope to inspire a new wave of research into peptideins and to unlock new insights and drug targets across human biology, particularly for the development of cellular immunotherapies and cancer vaccines.”
The study is the work of the TransCODE Consortium, an international collaboration of more than 60 researchers at over 30 institutions worldwide, co-led by the Princess Máxima Center for Pediatric Oncology in the Netherlands, the University of Michigan Medical School, the EMBL European Bioinformatics Institute in Hinxton, and the Institute for Systems Biology in Seattle.
Genes in DNA provide the recipe for cells to produce peptides. Historically, peptides have been called proteins if they are long enough and have existing evidence for a biological role, such as the appearance of the same protein across species in evolution. “Protein-coding genes are the bedrock of biomedical investigations, including the overwhelming majority of drug development programs,” the authors wrote. A large, curated international database of proteins contains some 19,500 entities.
But increasingly, scientists believe the traditional definition of a protein needs to be broadened. “Whether the human genome encodes substantially more than the approximately 19,500 canonical protein-coding genes has sparked a spirited debate in recent years,” the scientist continued. “Therefore, any wholesale addition of protein-coding genes creates ripple effects across human bioscience.”
Through their newly reported study the team looked at more than 7,200 previously understudied sections of the DNA called non-canonical open reading frames (ncORFs). They found that some 25% of these sections—more than 1,700—generated detectable protein-like molecules. These proteins, smaller than traditional proteins, are referred to as “microproteins.”
Generating their results involved looking at 3.7 billion individual bits of raw data that may support known and previously unknown proteins—drawing upon 95,520 experiments. “We show that about 25% of a set of 7,264 ncORFs gives rise to detectable peptides in a large-scale analysis of 95,520 proteomics experiments,” they wrote. The process took around 20,000 hours for computers to complete, working non-stop. They found 1,785 microproteins, a number that at first glance would increase the protein databases by nearly 10%.
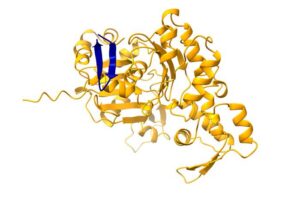
![Predicted binding between a non-canonical open reading frame (blue) and traditional protein (yellow). [Leron Kok/Princess Máxima Center for pediatric oncology]](https://www.genengnews.com/wp-content/uploads/2026/05/Low-Res_Protein-structure-copyright-Leron-Kok-Princess-Maxima-Center-300x201.jpg)
Moritz further explained, “By deploying our battle-hardened Trans Proteomic Pipeline across nearly 100,000 mass spectrometry experiments encompassing 3.7 billion spectra—derived from the world’s collective publicly available mass spectrometry data, with the results housed within PeptideAtlas at ISB for the scientific community to view and share—we were able to confirm, with high confidence, the existence of more than 1,700 of these newly identified peptideins that would otherwise have largely remained invisible to science.”
But most of these 1,785 microproteins didn’t resemble the other 19,500 traditional proteins. For example, they were very small: 65% were fewer than 50 amino acids in length, compared to less than 1% of the 19,500 previously catalogued. Looking more closely at the microproteins the investigators saw that only a few—perhaps a dozen—resembled the traditional proteins. The team then spent more than a year trying to make sense out of the remaining bulk.
Working with protein experts from across the globe in the TransCODE consortium, the scientists coined a new biological concept, which they coined peptidein. For decades, the research community has had a binary view of the relationship between human DNA and human proteins. A given piece of DNA either does or does not produce a protein. In their new study, the scientists propose a third choice, which is that DNA could make a protein, a peptidein, or neither.
The team defined a peptidein as existing in cells as a protein-like molecule, meaning that it is made of amino acids, as are proteins. But the role of a peptidein is ambiguous. Perhaps it has a function in normal human biology, or perhaps not; this is the key distinction with traditional proteins, where all are believed to have a function in normal human biology even if the details of that function are not fully known yet. “To advance these ncORFs in biological inquiry, we invoke the emerging umbrella term of peptidein, which we define as an ORF with experimentally confirmed RNA translation and protein synthesis, but for which the data are currently insufficient to claim conventional protein-coding gene status,” the investigators stated in their report.
Importantly, this definition of peptidein leaves the door open for it to become a ‘protein’ in the future—that is, if scientists gather more evidence on it. To start exploring this idea, the team searched for peptideins without which cells cannot survive. These so-called pan-essential peptideins can be important candidate drug targets in cancer and other diseases.
Using large-scale CRISPR gene editing, the scientists found six peptideins that looked promising. For example, one of these was a peptidein produced from OLMALINC, a genetic sequence previously thought not to produce proteins. When the researchers switched this gene off, 85% of more than 485 cancer cell lines showed impaired survival. The researchers confirmed that this effect comes from the peptidein itself, not the RNA molecule it sits on, and found that it plays a role in cell division and DNA damage response. “Our work here highlights c10riboseqorf92 (in the OLMALINC transcript),” they commented. “… while we do not yet have sufficient evidence that this ncORF encodes a bona fide protein, its CRISPR-based phenotypes in the context of cancer cells are intriguing.”
Many of the newly detected peptideins are presented on cell surfaces for recognition by the immune system, making them potential targets for cancer immunotherapy. A number of such molecules presented to the immune system are already under development as drug targets, and there is growing interest from both academia and industry in exploiting this new class of cancer antigens. Peptideins could also shed light on genetic diseases that conventional gene analysis has been unable to explain, simply because genetic diagnostics were unaware that these molecules were encoded by the human genome.
Members of the consortium had previously uncovered an essential role for a microprotein, ASNSD1-uORF, in children with a high-risk form of the brain cancer, medulloblastoma. Scientists at the Princess Máxima Center are now carrying out further research to determine its role in additional pediatric cancers with the activated MYC oncogene, such as neuroblastoma.
van Heesch commented, “It felt really special to discuss and decide what to do with this new class of molecules, as we had gathered enough early evidence to suspect that they might be widespread across cell types and tissues. By classifying these molecules of unknown functionality as peptideins, we’ve given them a formal place in reference databases so the wider community can study them.”
In their paper the researchers concluded, “The extent of the undiscovered proteome is one of the central questions in human biomedicine. This work reflects the multi-consortium collaboration between the TransCODE Consortium, the HUPO-HPP/PeptideAtlas project, the HIPP immunopeptidomics project and the GENCODE gene annotation group to coalesce a generalizable approach towards understanding which ncORFs can be understood as encoding proteins … Through our efforts, we bring microproteins and alternative protein molecules into reference gene annotation by defining them as either a protein-coding gene or a peptidein, a new concept referring to confirmed protein molecules of indeterminate consequence.”
Prensner added, “We’re just beginning to see what this ‘dark proteome’ has to offer. It’s like the trailer to a movie. We see the outline of a game-changing view of human biology. We’re incredibly excited that the coming years will open new doors to help solve and treat human diseases such as cancer.”
Moritz further stated, “Our collaborative work represents a culmination of decades of investment from federal funding agencies in building the computational and data infrastructure needed to interrogate the proteome at truly unprecedented scale at the Institute for Systems Biology … What excites me most is not simply that these molecules exist, but what their existence implies.”
The researchers are making we make all ncORFs, peptides and spectra publicly available through PeptideAtlas.
The post Microproteins and Peptideins Expand Boundaries of the Human Proteome appeared first on GEN – Genetic Engineering and Biotechnology News.
Uncategorized
Amgen, AbbVie say IRA negotiations impacted Q1 sales
At least two drugmakers say they’re feeling the impact of the Inflation Reduction Act, after the first drug prices negotiated under the law took effect in January.
Amgen and AbbVie reported in their first-quarter earnings …
-

 Uncategorized9 years ago
Uncategorized9 years agoThese ’90s fashion trends are making a comeback in 2017
-

 Contributors9 years ago
Contributors9 years agoThe final 6 ‘Game of Thrones’ episodes might feel like a full season
-

 Uncategorized9 years ago
Uncategorized9 years agoAccording to Dior Couture, this taboo fashion accessory is back
-

 Uncategorized9 years ago
Uncategorized9 years agoUber and Lyft are finally available in all of New York State
-

 Uncategorized9 years ago
Uncategorized9 years agoPhillies’ Aaron Altherr makes mind-boggling barehanded play
-

 Uncategorized9 years ago
Uncategorized9 years agoThe old and New Edition cast comes together to perform
-

 Uncategorized9 years ago
Uncategorized9 years agoSteph Curry finally got the contract he deserves from the Warriors
-

 Uncategorized9 years ago
Uncategorized9 years agoDisney’s live-action Aladdin finally finds its stars